In research and drug development contexts such as protein labeling, antibody conjugation, and the construction of multifunctional fusion proteins, site-specific protein ligation technologies are becoming increasingly important [1]. Among these, Sortase A (SrtA), a transpeptidase derived from Gram-positive bacteria, has become one of the most widely used bioconjugation tools due to its ability to specifically recognize the LPXTG motif and catalyze site-specific ligation with oligoglycine substrates. As application complexity continues to increase, researchers have further improved the catalytic efficiency and reaction performance of Sortase A through protein engineering and directed evolution, enabling more efficient and controllable protein conjugation.
1. What is Sortase A Protease?
Sortase A protease is a naturally occurring transpeptidase that recognizes and acts on specific short peptide sequences, most notably the LPXTG recognition motif [2,3]. Under mild reaction conditions, Sortase A identifies the LPXTG sequence within a protein and performs precise cleavage between the threonine (T) and glycine (G) residues. Here, X represents any amino acid and has minimal impact on overall recognition. The resulting enzyme–substrate intermediate can subsequently undergo a transpeptidation reaction with molecules bearing an N-terminal oligoglycine (typically GGG), enabling site-specific ligation of new peptide segments or functional modules at the defined position.
Unlike random chemical modification methods that rely on chemical activation, Sortase A-mediated reactions exhibit high sequence specificity, producing well-defined and predictable products. This makes it particularly suitable for protein systems that are highly sensitive to structural integrity and functional preservation.
Therefore, Sortase A serves as a transpeptidase tool capable of precise protein modification at predefined sites.
2. Application I: Use of Sortase A in Tag Removal
2.1 Why is enzymatic tag removal necessary?
During the expression and purification of recombinant proteins, various affinity tags (such as His-tag and Flag-tag) are indispensable tools. However, the presence of these tags is not the end goal. After obtaining a highly purified target protein, residual tags may interfere with downstream studies, for example:
- Altering the native conformation of the protein
- Affecting biological activity or binding capability
- Interfering with subsequent site-specific labeling, conjugation, or functional evaluation assays
Therefore, the safe and controllable removal of fusion tags during functional protein preparation is a critical step to ensure experimental quality.
2.2 Advantages of Sortase A as a tag removal tool
Using Sortase A for site-specific cleavage enables precise removal of tags at a pre-designed recognition sequence, offering the following advantages [4]:
- Defined cleavage site without introducing extra residues
- Mild reaction conditions, suitable for structurally and functionally sensitive proteins
- Highly predictable products, facilitating downstream quality control
Notably, the site-specific controllability demonstrated by Sortase A in tag removal also lays the foundation for its broader application in subsequent site-specific protein modification.
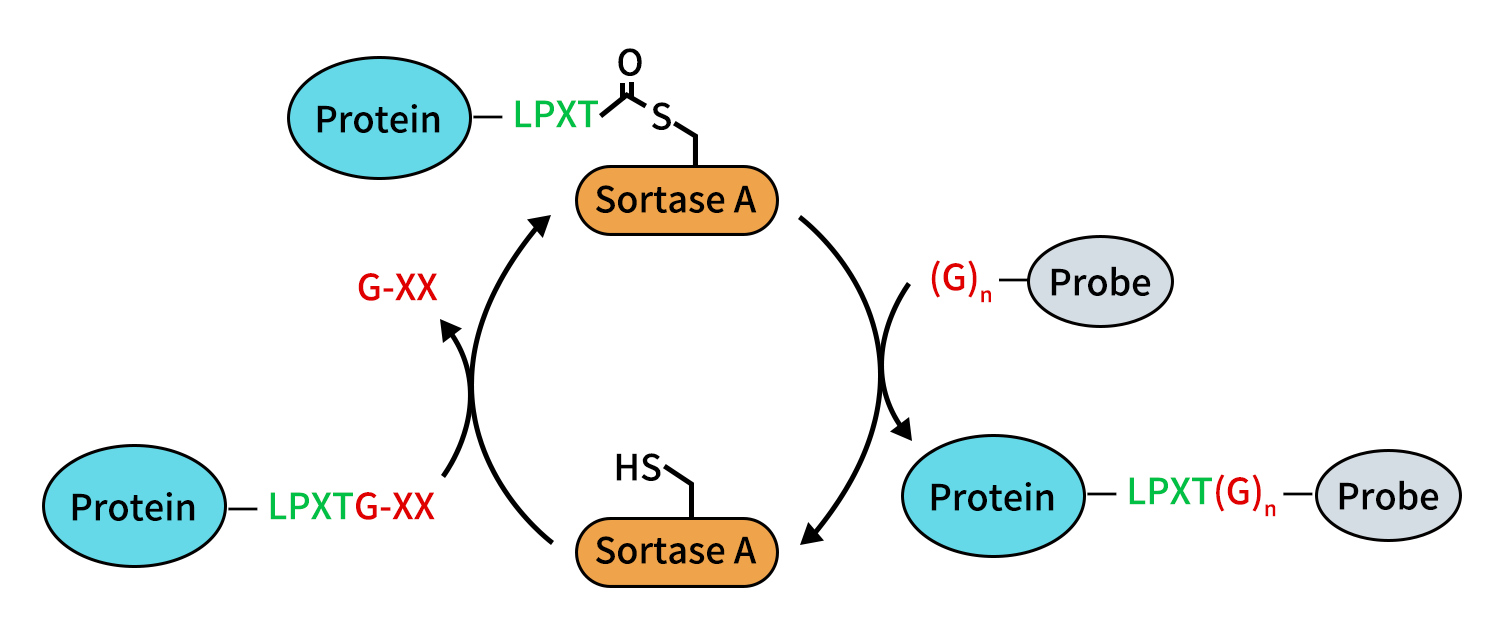
Figure 1. N-terminal labeling of proteins.
3. Application II: Sortase A–Mediated Site-Specific Protein Labeling and Conjugation
3.1 Why is site-specific labeling replacing random labeling?
In protein functional studies and analytical systems, the introduction of labeling molecules is often unavoidable. Compared with conventional random labeling, site-specific labeling offers significant advantages [5,6]:
- Avoids over-modification of key functional regions
- Significantly improves experimental reproducibility and data consistency
- Better suited for proteins that are highly sensitive to conformation and activity
As requirements for research precision continue to increase, site-specific labeling has become a preferred strategy in high-quality protein engineering projects.
3.2 Core advantages of Sortase A in site-specific labeling
Using a Sortase A–based site-specific conjugation system, researchers can introduce labeling molecules at predefined positions with uniform orientation. Its key advantages include [5–7]:
- Controllable labeling site and conjugation direction
- Highly uniform linkage, reducing sample heterogeneity
- Compatibility with bulky or structurally complex labeling molecules
3.3 Applications of Sortase A in antibody site-specific conjugation and ADC construction
In antibody-related conjugation applications, particularly antibody–drug conjugates (ADCs), the controllability of conjugation sites directly determines product homogeneity, stability, and in vivo behavior. Conventional random conjugation strategies based on lysine or cysteine residues often result in wide drug-to-antibody ratio (DAR) distributions and high structural heterogeneity, which can compromise efficacy and safety.
By introducing the LPXTG recognition motif at the C-terminus of the heavy or light chain of antibodies, Sortase A enables site-specific conjugation without interfering with antigen-binding regions. This allows the attachment of small-molecule cytotoxic drugs, linkers, fluorescent probes, or imaging tracers. This strategy yields well-defined conjugation sites and controllable DAR values, significantly improving batch-to-batch consistency and predictability of ADC molecules.
Currently, Sortase A-mediated antibody conjugation is widely used in ADC construction, mechanistic studies, and candidate screening, providing an important enabling tool for the development of next-generation homogeneous ADCs.
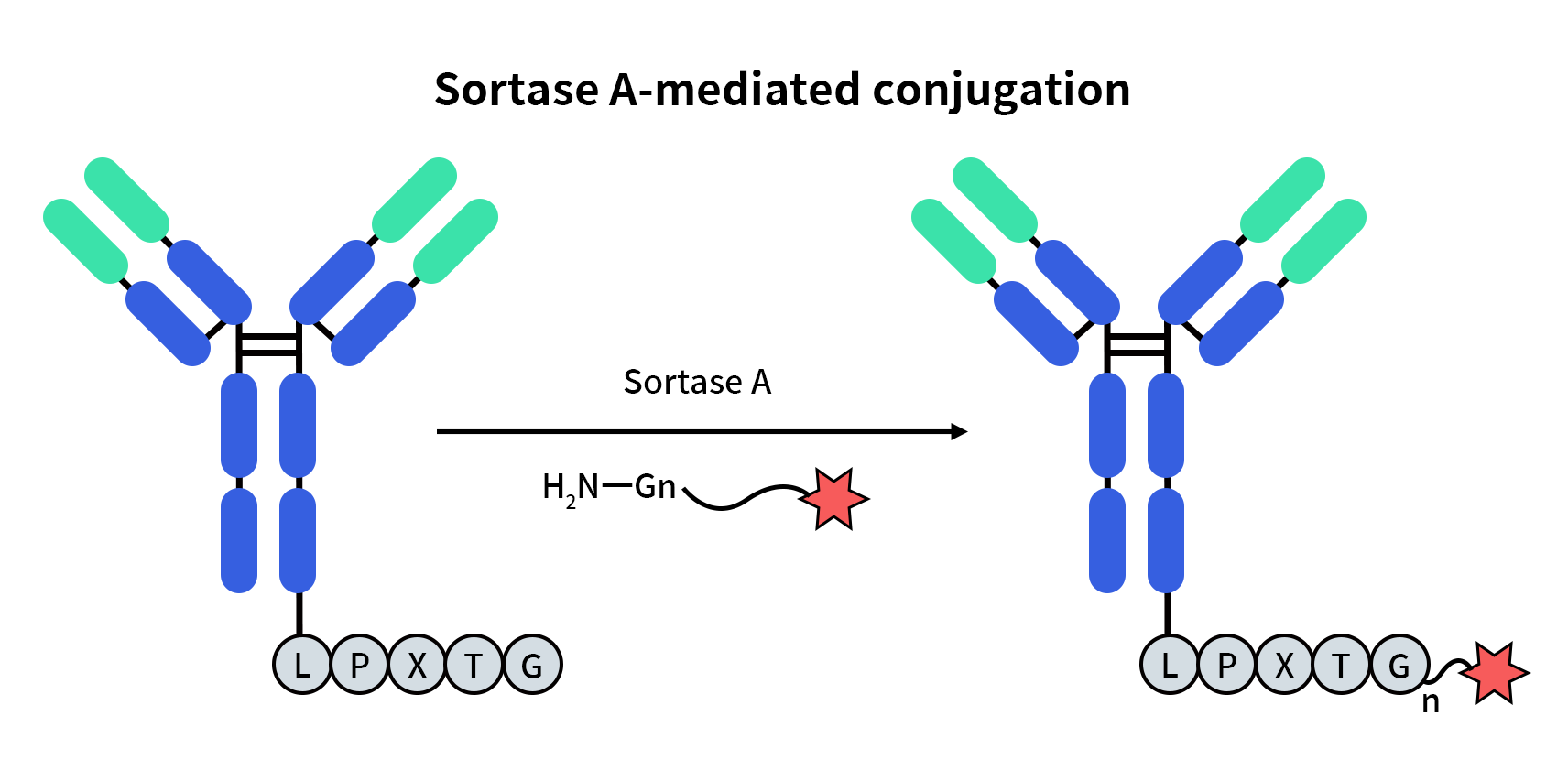
Figure 2. Sortase A-mediated conjugation.
4. Sortase A 5M: An Engineered Variant Optimized for Site-Specific Protein Engineering
As the demands for efficiency and system compatibility in site-specific protein engineering continue to increase, wild-type Sortase A has gradually revealed limitations in complex protein systems, including restricted reaction rates and insufficient substrate adaptability for large biomolecules. In particular, in applications involving antibodies, Fc-fusion proteins, and bulky labeling reagents, engineered variants with higher activity and improved stability have become critical for enhancing experimental robustness and success rates.
Sortase A 5M is a highly active engineered variant developed by Professor David R. Liu’s group through directed evolution. It contains five key mutations (P94R, D160N, D165A, K190E, and K196T) and retains strict specificity for the LPXTG recognition motif while significantly enhancing transpeptidation efficiency. Compared with the wild-type enzyme, this variant enables more efficient ligation under milder conditions and demonstrates improved substrate compatibility for Fc-fusion proteins, antibodies, and large functional tags [8]. In practical applications, its advantages are primarily reflected in the following aspects:
- First, higher transpeptidation efficiency, enabling stable and reproducible conjugation with lower enzyme usage and shorter reaction times.
- Second, improved reaction robustness, with greater tolerance to buffer conditions and substrate concentration variations, making it more suitable for standardized and scalable workflows.
- Third, enhanced compatibility with complex or large substrates, including Fc-fusion proteins, antibodies, and large fluorescent or functional labels.
The Sortase A 5M product provided by DIMA is based on this well-validated engineered enzyme system and can reliably support multiple applications, including tag removal, site-specific biotinylation, fluorescent protein conjugation, and antibody-related site-specific modifications. The biotin and PE labeling cases presented below are both established on the Sortase A 5M platform and have undergone internal functional validation.
5. DIMA Validation Applications Based on Sortase A 5M: Biotin and PE Labeling
5.1 Sortase A 5M–Mediated Site-Specific Biotin Labeling
In biotin labeling systems, random biotinylation often leads to impaired binding sites or unstable signals. Site-specific biotinylation using Sortase A 5M effectively avoids these issues. This strategy has been validated in internal applications and is compatible with multiple downstream analytical platforms, including ELISA, BLI/SPR kinetic analysis, and pull-down assays for interaction studies.
Case 1: BCMA-hFc fusion protein was selected as a model molecule. Using a Sortase A 5M–mediated C-terminal site-specific conjugation system, a biotin tag was introduced at its C-terminus. ELISA was then used to systematically evaluate antigen-binding activity retention before and after conjugation, thereby demonstrating the applicability and reliability of this strategy for functionalization of antibodies and immune-related proteins.
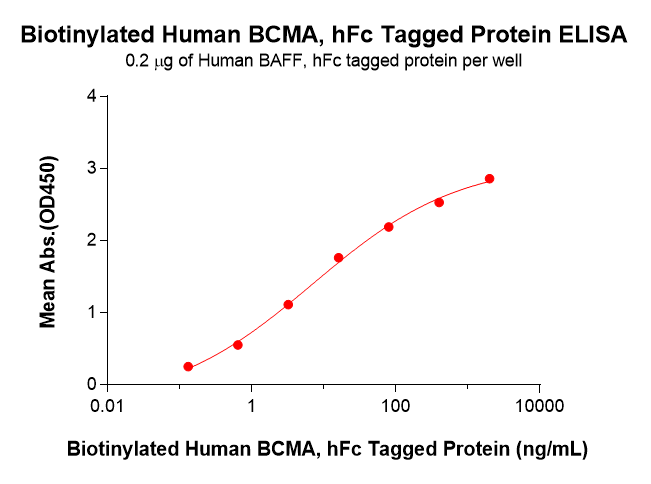
Figure 3. Biotinylated Human BCMA-hFc shows dose-dependent binding to immobilized Human BAFF-hFc, as detected by HRP–streptavidin, with an EC50 of 7.03 ng/mL.
Case 2: A monoclonal antibody against B7-H3 was used as a model system. Using a Sortase A 5M–mediated C-terminal site-specific biotinylation strategy, the antibody was precisely functionalized and subsequently applied in a sandwich ELISA system to evaluate the impact of this site-specific labeling approach on antibody detection performance and functional retention.
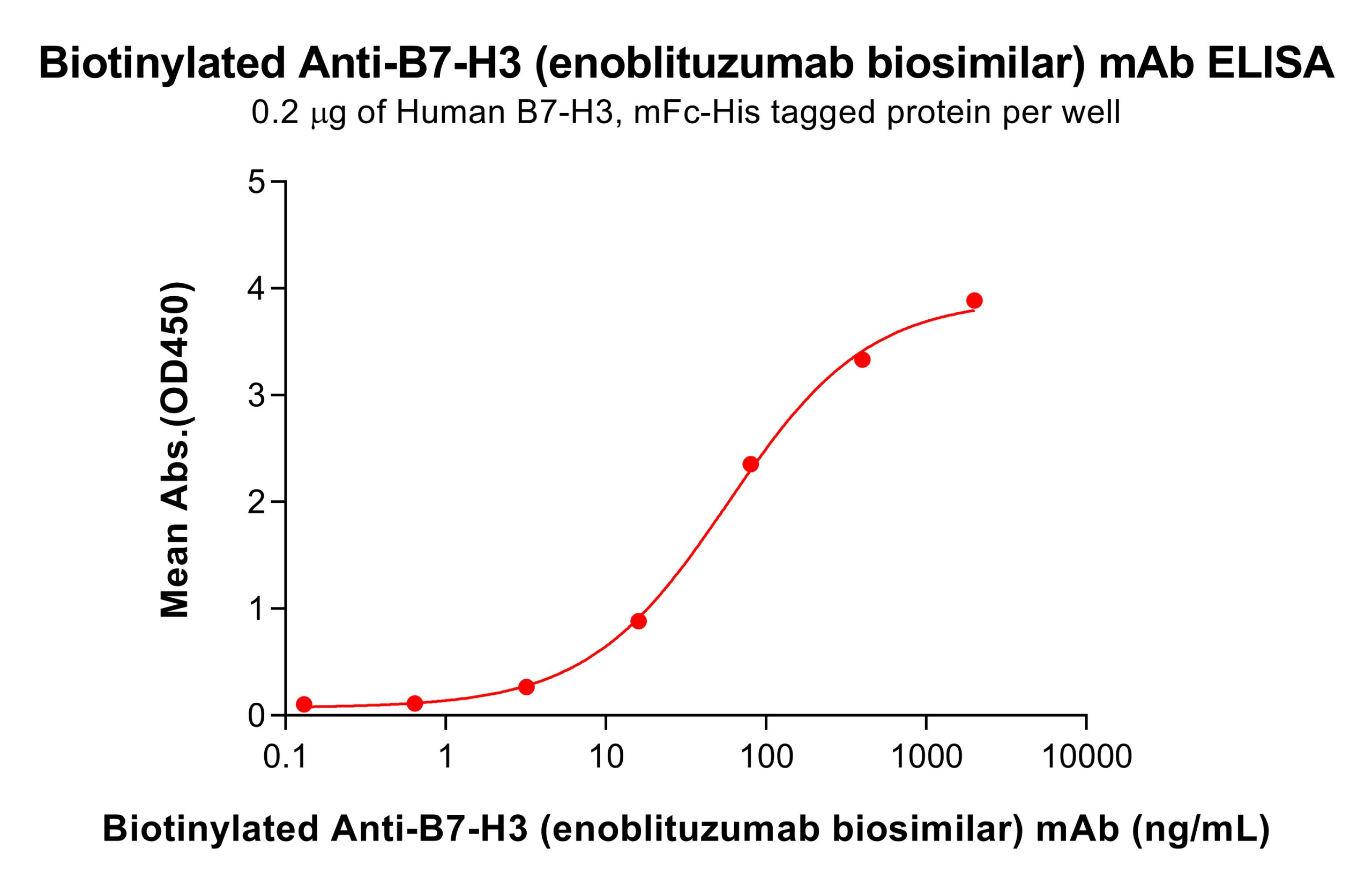
Figure 4. Biotinylated anti-B7-H3 mAb (enoblituzumab biosimilar) exhibits linear binding to ELISA-immobilized Human B7-H3 mFc-His within 16-80 ng/mL.
5.2 Sortase A 5M–Mediated Site-Specific PE Labeling
Phycoerythrin (PE) is a bulky fluorescent label that is highly sensitive to conjugation sites and spatial conformation. Random conjugation may not only reduce fluorescent performance but also disrupt the native structure of the protein. Site-specific conjugation of PE using Sortase A 5M enables linkage at defined positions with uniform orientation, preserving both fluorescence properties and protein structural integrity. This strategy has also been validated in internal applications and is particularly suitable for high-sensitivity detection platforms such as flow cytometry.
Case 1: To evaluate whether Sortase A 5M–mediated PE site-specific labeling is suitable for application scenarios that are highly sensitive to conformation and steric hindrance, such as flow cytometry, we further performed functional validation of the PE-labeled anti–B7-H3 antibody at the cellular level.
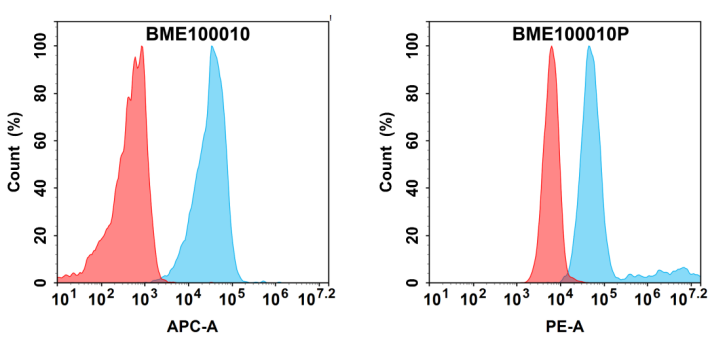
Figure 5. Flow cytometry comparison of Anti-B7-H3 (enoblituzumab biosimilar) mAb before and after PE conjugation on HEK293 cells. Blue: Anti-B7-H3; Red: isotype control.
6. Sortase A Is More Than a “Site-Specific Conjugation Enzyme”
From a technical standpoint, Sortase A is not a single-purpose conjugation tool, but rather a platform for site-specific protein engineering. It can be used both to remove undesired tags and to introduce functional modules or labeling moieties at defined positions. Its core value lies in its ability to precisely edit protein structure and function, providing a highly controllable strategy for the design and optimization of complex protein systems.
In summary, Sortase A is fundamentally a general-purpose enzymatic tool for site-specific protein engineering, rather than merely an auxiliary reagent for site-specific conjugation. To further support protein engineering research and functional applications, we provide high-quality Sortase A protein that can be broadly applied to tag removal, site-specific labeling, and targeted conjugation of antibodies or Fc-fusion proteins. Combined with our extensive validated application cases (e.g., BCMA-hFc, BM35 antibody, B7-H3 antibody, etc.), the use of this Sortase A protein can significantly improve experimental controllability, reproducibility, and data reliability.
Reference
- Popp, M. W., & Ploegh, H. L. (2011). Making and breaking peptide bonds: protein engineering using sortase. Angewandte Chemie International Edition, 50(22), 5024–5032.
- Mazmanian, S. K., Liu, G., Ton-That, H., & Schneewind, O. (1999). Staphylococcus aureus sortase, an enzyme that anchors surface proteins to the cell wall. Science, 285(5428), 760–763.
- Ton-That, H., Mazmanian, S. K., Alksne, L., & Schneewind, O. (2002). Anchoring of surface proteins to the cell wall of Staphylococcus aureus. Journal of Biological Chemistry, 277(9), 7447–7452.
- Witte, M. D., Cragnolini, J. J., Dougan, S. K., Yoder, N. C., Popp, M. W., & Ploegh, H. L. (2012). Preparation of unnatural N-to-N and C-to-C protein fusions. Proceedings of the National Academy of Sciences, 109(30), 11993–11998.
- Antos, J. M., Chew, G.-L., Guimaraes, C. P., Yoder, N. C., Grotenbreg, G. M., Popp, M. W., & Ploegh, H. L. (2009). Site-specific protein labeling via sortase-mediated transpeptidation. Current Protocols in Protein Science, Chapter 15, Unit 15.3.
- Theile, C. S., et al. (2013). Site-specific N-terminal labeling of proteins using sortase-mediated reactions. Nature Protocols, 8(9), 1800–1807.
- Guimaraes, C. P., Carette, J. E., Varadarajan, M., Antos, J., Popp, M. W., Spooner, E., Brummelkamp, T. R., & Ploegh, H. L. (2011). Identification of host cell factors required for intoxication through use of modified cholera toxin. The Journal of cell biology, 195(5), 751–764.
- Chen, B. M. Dorr & D. R. Liu, A general strategy for the evolution of bond-forming enzymes using yeast display, Proc. Natl. Acad. Sci. U.S.A., 108(28):11399–11404 (2011).
